
Sara Ayuso-Tejedor, Shingo Nishikori , Takashi Okuno , Teru Ogura* & Javier Sancho* (2010) FtsH cleavage of non-native conformations of proteins. J. Struc. Biol. May 8. [Epub ahead of print] *corresponding authors
FtsH is a peculiar prokaryotic protease with low unfoldase activity. Different reports have proposed that FtsH substrates could be either tagged proteins or proteins of low stability. We show here that FtsH degradation of 31 point mutants of Anabaena apoflavodoxin is inversely proportional to their conformational stabilities, and that the same applies to other substrate proteins. In contrast, highly stable proteins such as GST and holoflavodoxin are not degraded at all. Attempts to identify sequence tags signaling for degradation in apoflavodoxin fragments have been unsuccessful. Apoflavodoxin adopts three conformations: native, partly-unfolded and fully-unfolded. It is revealing that degradation of the 31 variants is proportional to the molar fraction of fully-unfolded molecules and inversely proportional to the fraction of stable apoflavodoxin molecules. This indicates that FtsH, rather than unfolding the protein, acts on the fraction that is already unfolded.
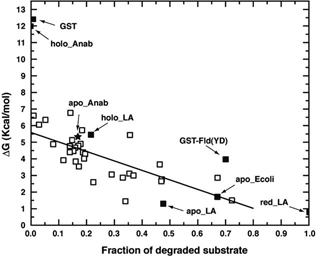
Figure FtsH degradation efficiency and substrate conformational stability. Degradation fractions by FtsH of wild-type Anabaena apoflavodoxin (closed star) and mutants (open squares) compared with their conformational stabilities. The degradation versus stability data corresponding to holo-lactalbumin (holo-LA), apo-LA, reduced (r)-LA, GST, apoflavodoxin from E. coli and holoflavodoxin from Anabaena are also shown as closed squares.